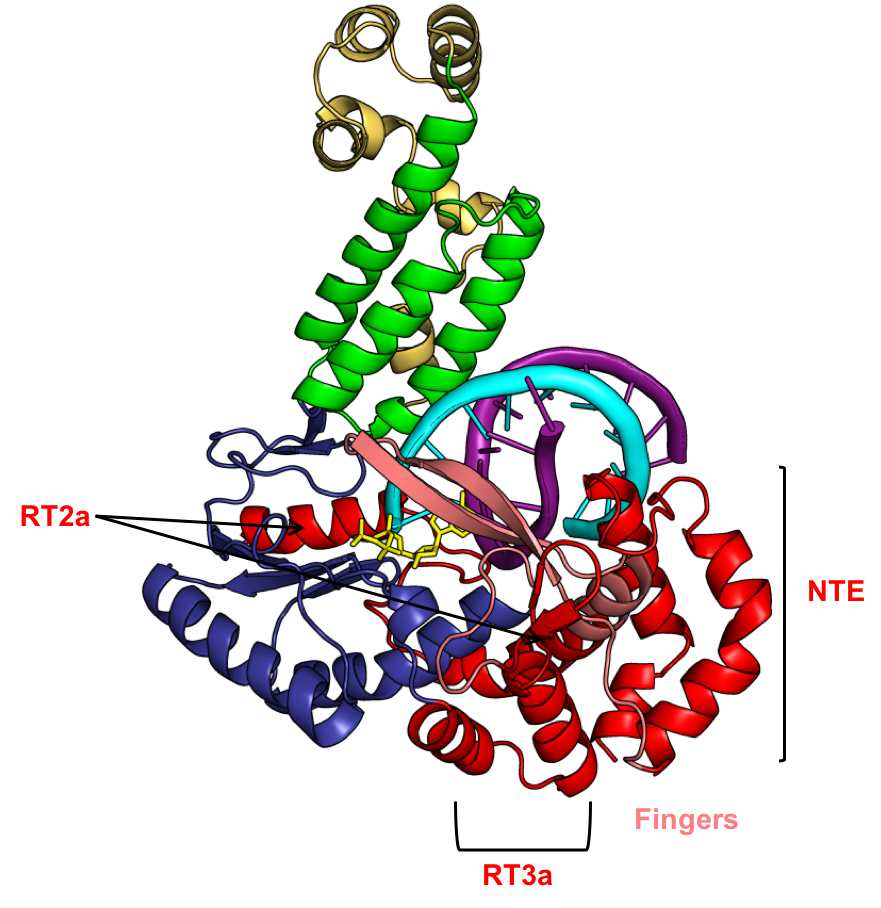
The research in our laboratory focuses on group II intron and related bacterial and organellar reverse transcriptases (RTs), their reverse transcription mechanisms, association with CRISPR-Cas systems, and applications in high-throughput RNA sequencing (RNA-seq), including for the analysis of human cellular, exosomal, and plasma RNAs for RNA diagnostics and liquid biopsy of human diseases. We use a combination of genetic, biochemical, microbiological, and structural approaches. Methods employed include X-ray crystallography and cryo-EM, as well as high-throughput RNA and DNA sequencing and a wide range of bioinformatic and computational methods for analyzing sequence data and biomarker identification in clinical samples, including in currently funded clinical trials. Current research topics are described under the "Research" tab.