Research
A grand challenge in molecular biology is to understand how biomolecules (e.g., proteins and RNA) reach their functional native states starting from an astronomical number of random coil conformations. The understanding of how these processes occur impacts the design of biomolecules of specified topology, and can offer critical clues to understanding diseases caused by conformational defects such as prions.
Our research group is primarily focused on discovering general principles that govern the folding of biomolecules. We use a combination of theoretical and computational strategies to answer several questions. In particular, we are interested in:
- Folding mechanisms of large protein and RNA molecules
- Causes of misfolded conformations in proteins
- Variations in the folding mechanisms of proteins in response to changes in environment
- Role of chaperones in aiding protein folding
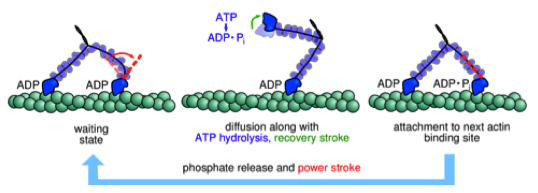
- Molecular basis for the function of molecular machines
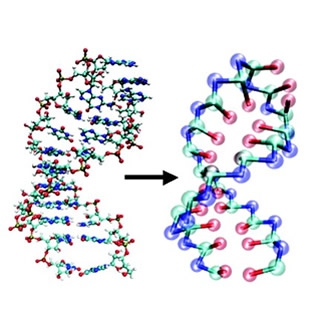
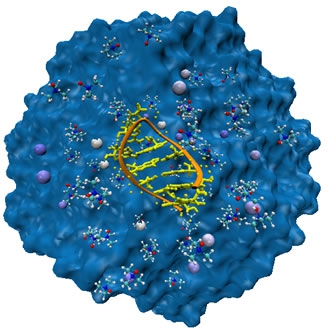
Another area of interest is in the study of changes in DNA shapes due to stretching and condensation. Currently, it is possible to manipulate single DNA molecules using optical tweezers. Such experiments provide direct measurements of the extension of DNA subject to force. The reversible stretching and condensation of DNA, which are important steps in the replication process, is being studied using statistical mechanical models. Such studies will provide detailed, microscopic description of the collapse of DNA and other synthetic polyelectrolytes and polyampholytes.

Molecular Motor Kinesin Walks Like a Drunk Man
Kinesin, an ATP-based cellular cargo transporter, takes a 16 nm step on microtubule.
GFP Folding Pathway (Govardhan Reddy)
